初めに
以下4つの次元削減アルゴリズムをPythonで実行し、それぞれで2次元のグラフを作成してみます。
次元圧縮したデータはmnistになります。実装や可視化をしたい方の参考になれば幸いです。
- PCA(Principal Component Analysis:主成分分析)
- t-SNE(t-distributed Stochastic Neighbor Embedding)
- UMAP(Uniform Manifold Approximation and Projection)
- VAE(Variational Auto-Encoder)
共通の実行環境として以下のコードを実行してください。
後は使用したい手法のコードを実行していただけると同じような結果が出ると思います。
from tensorflow import keras
from tensorflow.keras import layers
import tensorflow as tf
import matplotlib.pyplot as plt
from scipy.stats import norm
import numpy as np
from matplotlib.cm import get_cmap
cmap = get_cmap("tab10")PCA
#3分34秒
from sklearn.decomposition import PCA
(x_train, _), (x_test, y_test) = keras.datasets.mnist.load_data()
# normalize
x_train = x_train / 255
x_test = x_test / 255
# reshape
x_train = x_train.reshape(x_train.shape[0], -1)
x_test = x_test.reshape(x_test.shape[0], -1)
X_decomposed = PCA(n_components=2).fit_transform(x_test)
fig, ax = plt.subplots(1, 1, figsize=(12, 8))
cmap = get_cmap("tab10")
for i, y in enumerate(y_test):
marker = "$" + str(y) + "$"
idx = i
ax.scatter(X_decomposed[idx][0], X_decomposed[idx][1],marker=marker,color=cmap(y))
ax.set_title("PCA")
fig.savefig('./PCA.png')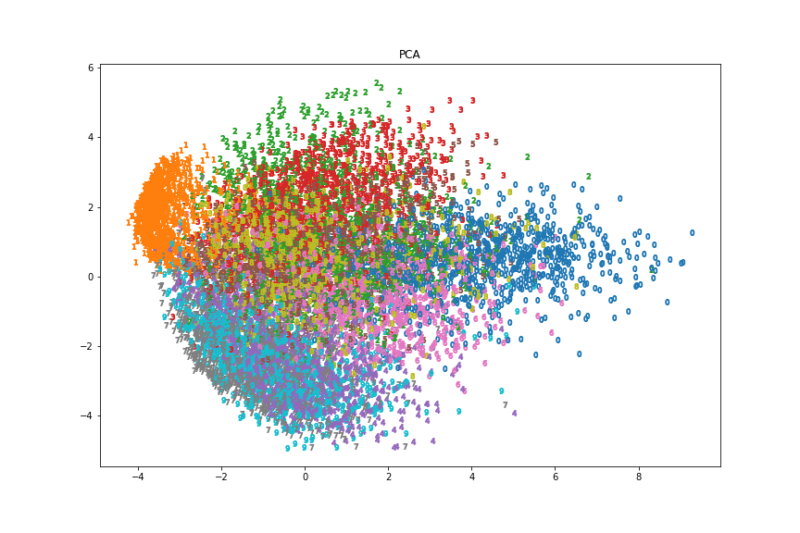
t-SNE
#6分11秒
from sklearn.manifold import TSNE
(x_train, _), (x_test, y_test) = keras.datasets.mnist.load_data()
# normalize
x_train = x_train / 255
x_test = x_test / 255
# reshape
x_train = x_train.reshape(x_train.shape[0], -1)
x_test = x_test.reshape(x_test.shape[0], -1)
X_decomposed = TSNE(n_components=2).fit_transform(x_test)
fig, ax = plt.subplots(1, 1, figsize=(12, 8))
cmap = get_cmap("tab10")
for i, y in enumerate(y_test):
marker = "$" + str(y) + "$"
idx = i
ax.scatter(X_decomposed[idx][0], X_decomposed[idx][1],marker=marker,color=cmap(y))
ax.set_title("t-SNE")
fig.savefig('./t-SNE.png')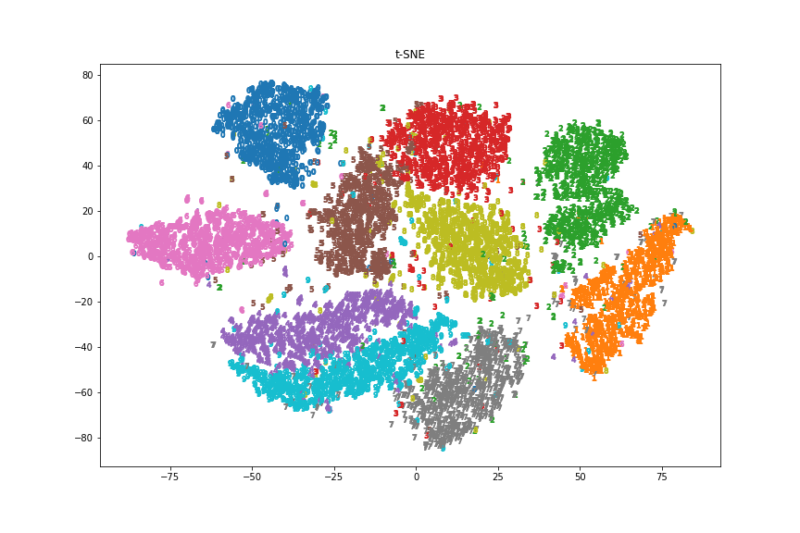
UMAP
#4分30秒
import umap
(x_train, _), (x_test, y_test) = keras.datasets.mnist.load_data()
# normalize
x_train = x_train / 255
x_test = x_test / 255
# reshape
x_train = x_train.reshape(x_train.shape[0], -1)
x_test = x_test.reshape(x_test.shape[0], -1)
mapper = umap.UMAP(random_state=0)
embedding = mapper.fit_transform(x_test)
fig, ax = plt.subplots(1, 1, figsize=(12, 8))
# 結果を二次元でプロットする
for i, y in enumerate(y_test):
marker = "$" + str(y) + "$"
idx = i
ax.scatter(embedding[idx][0], embedding[idx][1],marker=marker,color=cmap(y))
ax.set_title("UMAP")
fig.savefig('./umap.png')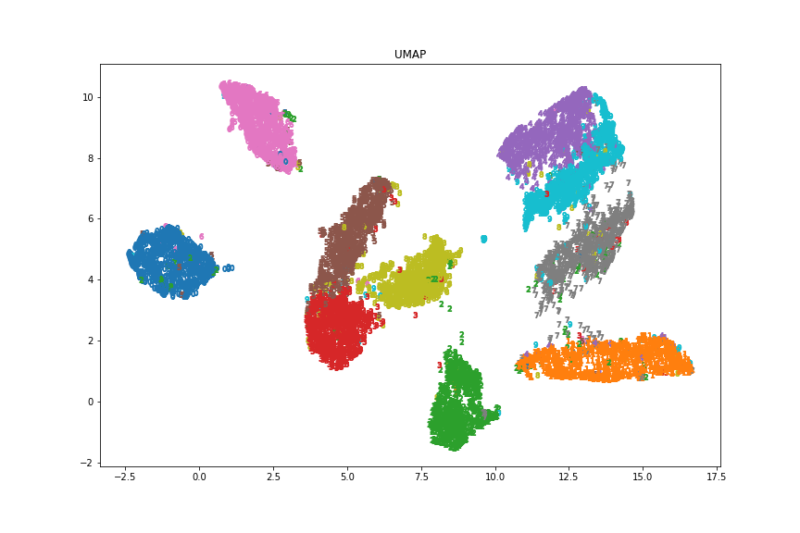
VAE
VAEに関しては以下のコードを順に実行してみてください。
encoder network
latent_dim = 2
encoder_inputs = keras.Input(shape=(28, 28, 1))
x = layers.Conv2D(32, 3, activation="relu", strides=2, padding="same")(encoder_inputs)
x = layers.Conv2D(64, 3, activation="relu", strides=2, padding="same")(x)
x = layers.Flatten()(x)
x = layers.Dense(16, activation="relu")(x)
z_mean = layers.Dense(latent_dim, name="z_mean")(x)
z_log_var = layers.Dense(latent_dim, name="z_log_var")(x)
encoder = keras.Model(encoder_inputs, [z_mean, z_log_var], name="encoder")潜在空間のサンプリング方法
https://jun2life.com/2021/11/11/ae%e3%81%a8vae%e3%81%ab%e3%81%a4%e3%81%84%e3%81%a6/
class Sampler(layers.Layer):
def call(self, z_mean, z_log_var):
batch_size = tf.shape(z_mean)[0]
z_size = tf.shape(z_mean)[1]
epsilon = tf.random.normal(shape=(batch_size, z_size))
return z_mean + tf.exp(0.5 * z_log_var) * epsilondecoder network
latent_inputs = keras.Input(shape=(latent_dim,))
x = layers.Dense(7 * 7 * 64, activation="relu")(latent_inputs)
x = layers.Reshape((7, 7, 64))(x)
x = layers.Conv2DTranspose(64, 3, activation="relu", strides=2, padding="same")(x)
x = layers.Conv2DTranspose(32, 3, activation="relu", strides=2, padding="same")(x)
decoder_outputs = layers.Conv2D(1, 3, activation="sigmoid", padding="same")(x)
decoder = keras.Model(latent_inputs, decoder_outputs, name="decoder")VAEモデル
class VAE(keras.Model):
def __init__(self, encoder, decoder, **kwargs):
super().__init__(**kwargs)
self.encoder = encoder
self.decoder = decoder
self.sampler = Sampler()
self.total_loss_tracker = keras.metrics.Mean(name="total_loss")
self.reconstruction_loss_tracker = keras.metrics.Mean(
name="reconstruction_loss")
self.kl_loss_tracker = keras.metrics.Mean(name="kl_loss")
@property
def metrics(self):
return [self.total_loss_tracker,
self.reconstruction_loss_tracker,
self.kl_loss_tracker]
def train_step(self, data):
with tf.GradientTape() as tape:
z_mean, z_log_var = self.encoder(data)
z = self.sampler(z_mean, z_log_var)
reconstruction = decoder(z)
reconstruction_loss = tf.reduce_mean(
tf.reduce_sum(
keras.losses.binary_crossentropy(data, reconstruction),
axis=(1, 2)
)
)
kl_loss = -0.5 * (1 + z_log_var - tf.square(z_mean) - tf.exp(z_log_var))
total_loss = reconstruction_loss + tf.reduce_mean(kl_loss)
grads = tape.gradient(total_loss, self.trainable_weights)
self.optimizer.apply_gradients(zip(grads, self.trainable_weights))
self.total_loss_tracker.update_state(total_loss)
self.reconstruction_loss_tracker.update_state(reconstruction_loss)
self.kl_loss_tracker.update_state(kl_loss)
return {
"total_loss": self.total_loss_tracker.result(),
"reconstruction_loss": self.reconstruction_loss_tracker.result(),
"kl_loss": self.kl_loss_tracker.result(),
}VAEモデルの学習
#10分
(x_train, _), (x_test, _) = keras.datasets.mnist.load_data()
mnist_digits = np.concatenate([x_train], axis=0)
mnist_digits = np.expand_dims(mnist_digits, -1).astype("float32") / 255
vae = VAE(encoder, decoder)
vae.compile(optimizer=keras.optimizers.Adam(), run_eagerly=True)
vae.fit(mnist_digits, epochs=30, batch_size=128)潜在空間からの画像マッピング
import matplotlib.pyplot as plt
n = 30
digit_size = 28
figure = np.zeros((digit_size * n, digit_size * n))
grid_x = np.linspace(-2, 2, n)
grid_y = np.linspace(-2, 2, n)[::-1]
for i, yi in enumerate(grid_y):
for j, xi in enumerate(grid_x):
z_sample = np.array([[xi, yi]])
x_decoded = vae.decoder.predict(z_sample)
digit = x_decoded[0].reshape(digit_size, digit_size)
figure[
i * digit_size : (i + 1) * digit_size,
j * digit_size : (j + 1) * digit_size,
] = digit
plt.figure(figsize=(15, 15))
start_range = digit_size // 2
end_range = n * digit_size + start_range
pixel_range = np.arange(start_range, end_range, digit_size)
sample_range_x = np.round(grid_x, 1)
sample_range_y = np.round(grid_y, 1)
plt.xticks(pixel_range, sample_range_x)
plt.yticks(pixel_range, sample_range_y)
plt.xlabel("z[0]")
plt.ylabel("z[1]")
plt.axis("off")
plt.imshow(figure, cmap="Greys_r")
plt.savefig('./VAE_plot.png')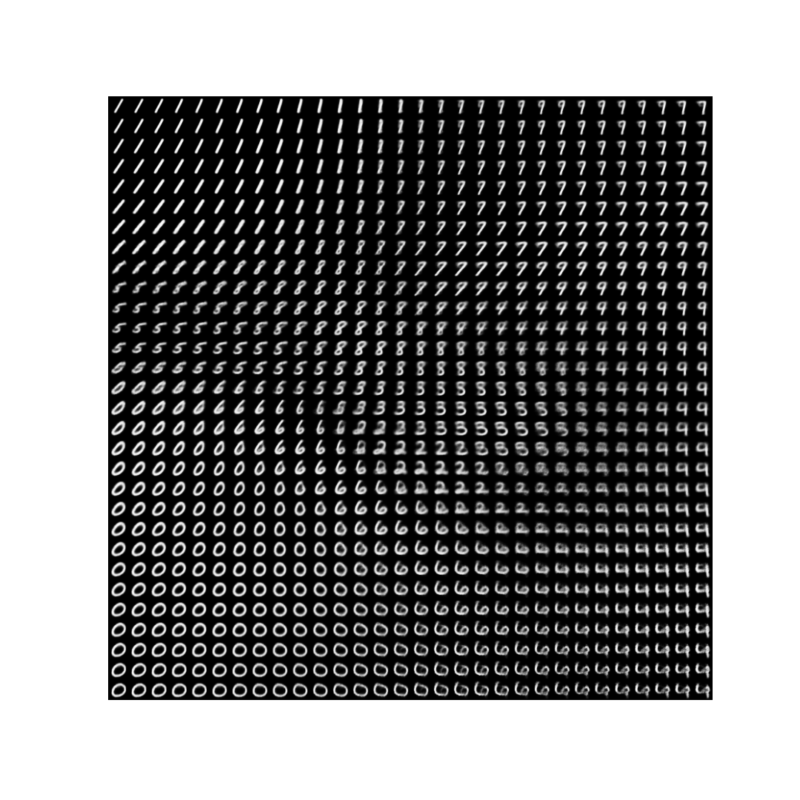
(x_train, _), (x_test, y_test) = keras.datasets.mnist.load_data()
test_digits = np.concatenate([x_test], axis=0)
test_digits = np.expand_dims(test_digits, -1).astype("float32") / 255
X_encoded = vae.encoder.predict(test_digits, batch_size=1)
# plot
fig, ax = plt.subplots(1, 1, figsize=(12, 8))
for i, y in enumerate(y_test):
marker = "$" + str(y) + "$"
idx = i
vec = X_encoded[0][idx] + np.exp(X_encoded[1][idx])
ax.scatter(vec[0], vec[1],marker=marker,color=cmap(y))
ax.set_title("VAE's latent space")
fig.savefig('./VAE.png')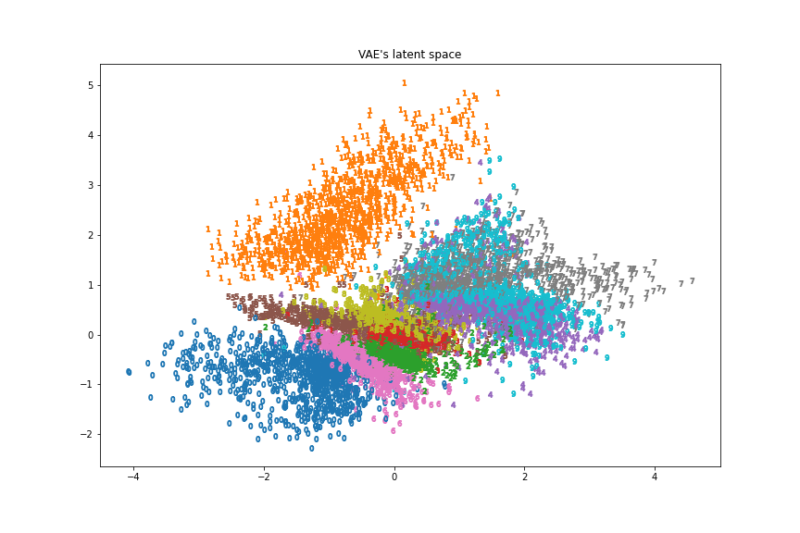
まとめ
画像と実行速度を見るとt-SNEが一番良さそうでした。



コメント